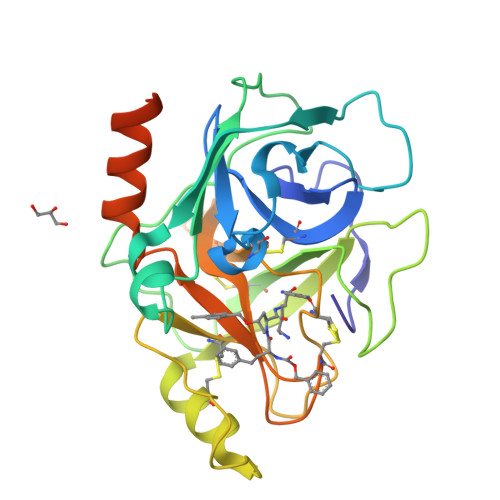
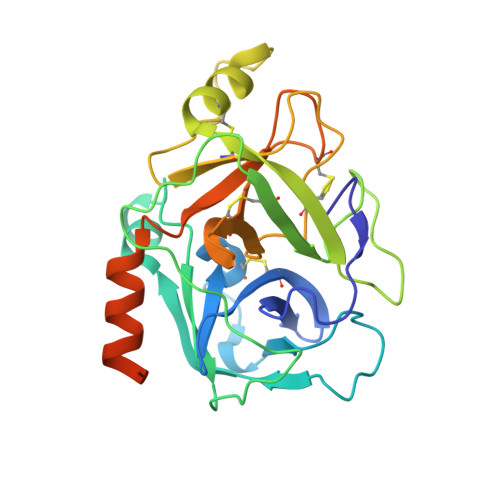
Discovery of inhibitors of the channel-activating protease prostasin (CAP1/PRSS8) utilizing structure-based design.
Tully, D.C., Vidal, A., Chatterjee, A.K., Williams, J.A., Roberts, M.J., Petrassi, H.M., Spraggon, G., Bursulaya, B., Pacoma, R., Shipway, A., Schumacher, A.M., Danahay, H., Harris, J.L.(2008) Bioorg Med Chem Lett 18: 5895-5899
- PubMed: 18752942
- DOI: https://doi.org/10.1016/j.bmcl.2008.08.029
- Primary Citation of Related Structures:
3E0P, 3E16 - PubMed Abstract:
Structure-based design was utilized to guide the early stage optimization of a substrate-like inhibitor to afford potent peptidomimetic inhibitors of the channel-activating protease prostasin. The first X-ray crystal structures of prostasin with small molecule inhibitors bound to the active site are also reported.
Organizational Affiliation:
Genomics Institute of the Novartis Research Foundation, 10675 John J. Hopkins Dr., San Diego, CA 92121, USA. dtully@gnf.org